Pushing Genomic Science Forward with AI Acceleration
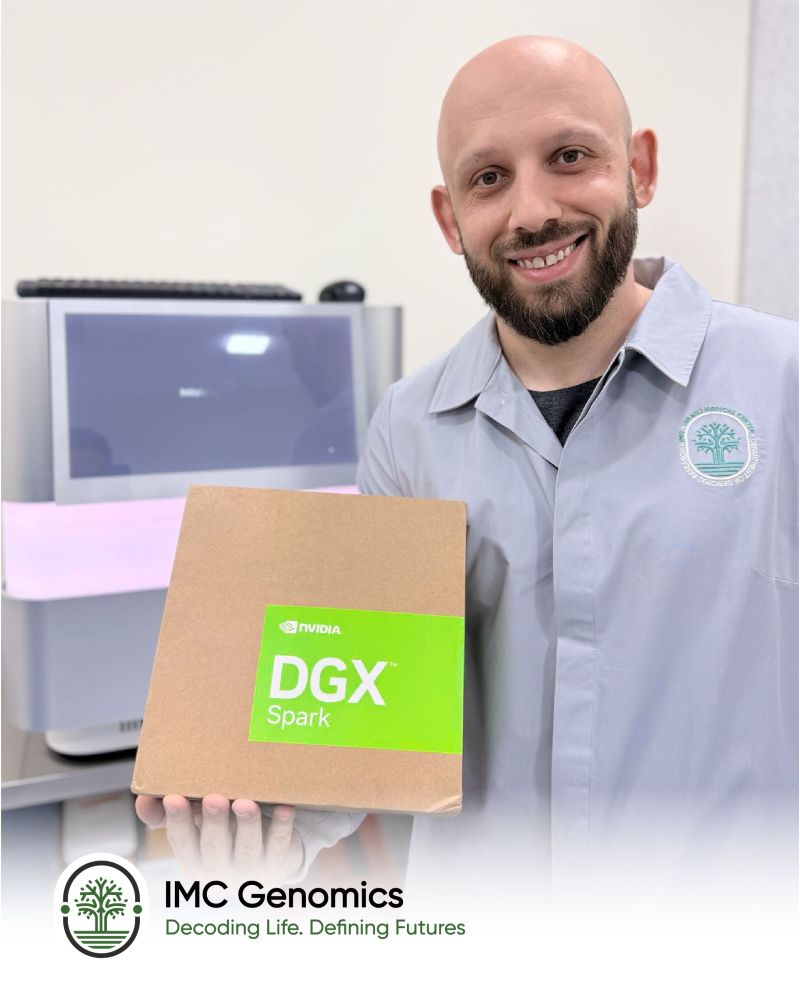

We've received the NVIDIA DGX Spark, marking another step toward scaling our computational genomics capabilities with AI-driven analytics.
IMC Genomics has taken a major step toward scaling our computational genomics pipeline with the arrival of the NVIDIA DGX Spark — a purpose-built AI workstation designed for data-intensive scientific workflows.
As next-generation sequencing generates terabytes of raw data per run, the challenge is no longer just sequencing — it is interpreting that data accurately and quickly. Traditional computing infrastructure can take hours to process a single whole-exome dataset. With the DGX Spark's dedicated GPU architecture, we can compress that timeline dramatically, enabling faster variant calling, annotation, and classification.
For our laboratory, this means several practical improvements. Turnaround times for complex panels can be reduced by accelerating the bioinformatics pipeline that transforms raw sequencing reads into clinically actionable reports. Machine learning models for variant pathogenicity prediction can be trained and refined locally rather than relying solely on cloud resources, giving us greater control over data security and pipeline customization.
The DGX Spark also positions us to explore emerging areas of computational genomics. Polygenic risk scoring, for example, requires processing thousands of genetic variants simultaneously across reference populations — a task that benefits enormously from parallel GPU computation. Similarly, AI-assisted quality control for sequencing runs can flag anomalies in real time, reducing the need for costly re-runs.
Central Asia's genomic infrastructure is still developing, and having this level of compute power in a private laboratory is uncommon in the region. We see it as part of our broader commitment to bringing world-class genomic capabilities closer to the patients and clinicians who need them — without requiring samples to be shipped overseas for analysis.
The DGX Spark joins our existing infrastructure alongside the Illumina NextSeq 2000, Thermo Fisher Ion Torrent systems, and our clinical bioinformatics pipeline, creating an end-to-end workflow from sample to report that is both fast and locally controlled.